From a bacterium, surprisingly.
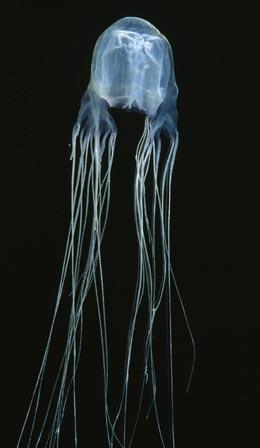 The venom of the box jellyfish can paralyse the central nervous system of its victims.NHPA/A.N.T. PHOTO LIBRARY
The venom of the box jellyfish can paralyse the central nervous system of its victims.NHPA/A.N.T. PHOTO LIBRARYJellyfish may owe thanks to a humble bacterium for their ability to sting prey. Scientists have found that one of the genes necessary for them to sting is similar to a gene in bacteria, suggesting the ancestors of jellyfish picked up the gene from microbes. The research is published this week in Current Biology1.
"The result was a great surprise," says developmental biologist Nicolas Rabet of the Pierre and Marie Curie University in Paris, France, who led the team. "[This kind of] horizontal gene transfer is often neglected, and could sometimes be more important than we thought." Unlike vertical gene transfer from parent to offspring, the horizontal variety happens between organisms, or even between different species. Common in microbes, it has only been described a few times in animals. Japanese beetles have picked up sequences from a parasitic bacterium2 and microscopic aquatic creatures called bdelloid rotifers have collected genes from bacteria, fungi and plants3.
The gene in question codes for a subunit of poly-gamma-glutamate (PGA) synthase - PGA itself is a major component of stinging cells4. The gene appears in all known genomes of creatures from the phylum cnidaria, which includes jellyfish, anemones and corals.
By collecting positive ions, PGA allows the cells to regulate their osmotic pressure; a sudden change in that pressure launches a poisonous barb. In bacteria, the same compound can form a protective capsule. It also gives the fermented Japanese food natto its stringy texture and pungent aroma.5
Using phylogenetic analysis, Rabet and his colleagues found that the cnidarian gene fits well into the bacterial family tree. They also showed that the gene turns on in at least one jellyfish, Clytia hemisphaerica. The same gene pops up in certain sponges, worms and fungi, suggesting it jumped between species more than once, the scientists say. It is not yet clear how the transfer might have occurred, or why this particular gene would be so well-travelled.
Vertical or horizontal?
"I think the author's interpretation is probably correct," says Michael Syvanen, who studies comparative genomics at the University of California, Davis. However, he is not convinced that other possibilites can be ruled out.
"There are other explanations for the incongruencies they see in the tree," agrees Casey Dunn, an evolutionary biologist who studies phylogenetic problems at Brown University in Providence, Rhode Island.
For instance, the gene could be vertically transferred from a distant progenitor, before being lost from some organisms. Or, it may be possible that more than one animal independently evolved the gene; such sequence conversion is not unheard of, Dunn says. "At the end of the day, it will probably take far more data to paint a conclusive picture of what's happening."
Rabet responds that since the PGA synthase gene is approximately 1000 bases long, it is statistically unlikely to be the product of multiple distinct genes converging on the same sequence
And if the gene was lost from all but the cnidarians and a few other animals, it must have disappeared from all related organisms. "It's possible, but we need to imagine a lot of lost genes," he says.
Scientists are finding that horizontal gene transfer, once thought to be the domain of single-celled critters, is not uncommon in the animal world, says Syvanen. "Horizontal gene transfer with the animals is going to turn out to be more widespread than anybody believes now. When that realization comes down, it will definitely change the way people think about evolution."





No comments:
Post a Comment